Edmonds's matching algorithm
Edmonds's matching algorithm [1] is an algorithm in graph theory for constructing maximum matchings on graphs. The algorithm was discovered by Jack Edmonds in 1965. Given a general graph G = (V, E), the algorithm finds a matching M such that each vertex in V is incident with at most one edge in M and |M| is maximized. The matching is constructed by iteratively improving an initial empty matching along augmenting paths in the graph. To search for augmenting paths, some odd-length cycles in the graph (blossoms) are contracted to single vertices and the search continues recursively in the contracted graphs.
Contents |
Augmenting paths
Given G = (V, E) and a matching M of G, a vertex v is exposed, if no edge of M is incident with v. A path in G is an alternating path, if its edges are alternately not in M and in M (or in M and not in M). An augmenting path P is an alternating path that starts and ends at two distinct exposed vertices. A matching augmentation along an augmenting path P is the operation of replacing M with a new matching 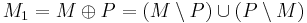 .
.
The algorithm uses the following fact:
Here is the high-level algorithm.
INPUT: Graph G, initial matching M on G OUTPUT: maximum matching M* on G A1 function find_maximum_matching( G, M ) : M* A2 P ← find_augmenting_path( G, M ) A3 if P is non-empty then A4 return find_maximum_matching( G, augment M along P ) A5 else A6 return M A7 end if A8 end function
The subroutine to find an augmenting path uses blossoms and contractions.
Blossoms and contractions
Given G = (V, E) and a matching M of G, a blossom B is a cycle in G consisting of 2k + 1 edges of which exactly k belong to M. We use G’, the contracted graph, to denote the graph obtained from G by contracting every edge of B. We use M’, the contracted matching, to denote the corresponding matching of G’. If P’ is a M’-augmenting path in G’ then P’ can be lifted to a M-augmenting path in G by undoing the contraction by B so that the segment of P’ (if any) traversing through vB is replaced by an appropriate segment traversing through B. In more detail:
- if P’ traverses through a segment u → vB → w in G’, then this segment is replaced with the segment u → ( u’ → ... → w’ ) → w in G, where blossom vertices u’ and w’ and the side of B, ( u’ → ... → w’ ), going from u’ to w’ are chosen to ensure that the new path is still alternating (u’ is exposed with respect to
 ,
, 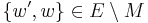 ).
).
- if P’ has an endpoint vB, then the path segment u → vB in G’ is replaced with the segment u → ( u’ → ... → v’ ) in G, where blossom vertices u’ and v’ and the side of B, ( u’ → ... → v’ ), going from u’ to v’ are chosen to ensure that the path is alternating (v’ is exposed,
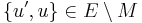 ).
).
The algorithm uses the following fact (using the notations from above):
- If B is a blossom, then G’ contains a M’-augmenting path if and only if G contains a M-augmenting path.[4] In addition, M’-augmenting paths in G’ correspond to M-augmenting paths in G.
Thus blossoms can be contracted and search performed in the contracted graphs. This reduction is at the heart of Edmonds's algorithm.
Finding an augmenting path
The search for augmenting path uses an auxiliary data structure consisting of a forest F whose individual trees correspond to specific portions of the graph G. Using the structure, the algorithm either (1) finds an augmenting path or (2) finds a blossom and recurses onto the corresponding contracted graph or (3) concludes there are no augmenting paths. The auxiliary structure is built by an incremental procedure discussed next.[4]
The construction procedure considers vertices v and edges e in G and incrementally updates F as appropriate. If v is in a tree T of the forest, we let root(v) denote the root of T. If both u and v are in the same tree T in F, we let distance(u,v) denote the length of the unique path from u to v in T.
INPUT: Graph G, matching M on G
OUTPUT: augmenting path P in G or empty path if none found
B01 function find_augmenting_path( G, M ) : P
B02 F ← empty forest
B03 unmark all vertices and edges in G, mark all edges of M
B05 for each exposed vertex v do
B06 create a singleton tree { v } and add the tree to F
B07 end for
B08 while there is an unmarked vertex v in F with distance( v, root( v ) ) even do
B09 while there exists an unmarked edge e = { v, w } do
B10 if w is not in F then
// Update F.
B11 x ← vertex matched to w in M
B12 add edges { v, w } and { w, x } to the tree of v
B13 else
B14 if distance( w, root( w ) ) is odd then
B15 do nothing
B16 else
B17 if root( v ) ≠ root( w ) then
// Report an augmenting path in F 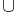 { e }.
B18 P ← path ( root( v ) → ... → v ) → ( w → ... → root( w ) )
B19 return P
B20 else
// Contract a blossom in G and look for the path in the contracted graph.
B21 B ← blossom formed by e and edges on the path v → w in T
B22 G’, M’ ← contract G and M by B
B23 P’ ← find_augmenting_path( G’, M’ )
B24 P ← lift P’ to G
B25 return P
B26 end if
B27 mark edge e
B28 end while
B29 mark vertex v
B30 end while
B31 return empty path
B32 end function
{ e }.
B18 P ← path ( root( v ) → ... → v ) → ( w → ... → root( w ) )
B19 return P
B20 else
// Contract a blossom in G and look for the path in the contracted graph.
B21 B ← blossom formed by e and edges on the path v → w in T
B22 G’, M’ ← contract G and M by B
B23 P’ ← find_augmenting_path( G’, M’ )
B24 P ← lift P’ to G
B25 return P
B26 end if
B27 mark edge e
B28 end while
B29 mark vertex v
B30 end while
B31 return empty path
B32 end function
Examples
The following four figures illustrate the execution of the algorithm. We use dashed lines to indicate edges that are currently not present in the forest. First, the algorithm processes an out-of-forest edge that causes the expansion of the current forest (lines B10 – B12).
Next, it detects a blossom and contracts the graph (lines B20 – B22).
Finally, it locates an augmenting path P′ in the contracted graph (line B23) and lifts it to the original graph (line B24). Note that the ability of the algorithm to contract blossoms is crucial here; the algorithm can not find P in the original graph directly because only out-of-forest edges between vertices at even distances from the roots are considered on line B17 of the algorithm.
Analysis
The forest F constructed by the find_augmenting_path() function is an alternating forest, .[5]
- a tree T in G is an alternating tree with respect to M, if
- T contains exactly one exposed vertex r called the tree root
- every vertex at an odd distance from the root has exactly two incident edges in T, and
- all paths from r to leaves in T have even lengths, their odd edges are not in M and their even edges are in M.
- a forest F in G is an alternating forest with respect to M, if
- its connected components are alternating trees, and
- every exposed vertex in G is a root of an alternating tree in F.
Each iteration of the loop starting at line B09 either adds to a tree T in F (line B10) or finds an augmenting path (line B17) or finds a blossom (line B21). It is easy to see that the running time is 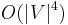 . Micali and Vazirani [6] show an algorithm that constructs maximum matching in
. Micali and Vazirani [6] show an algorithm that constructs maximum matching in 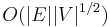 time.
time.
Bipartite matching
The algorithm reduces to the standard algorithm for matching in bipartite graphs [3] when G is bipartite. As there are no odd cycles in G in that case, blossoms will never be found and one can simply remove lines B21 – B29 of the algorithm.
Weighted matching
The matching problem can be generalized by assigning weights to edges in G and asking for a set M that produces a matching of maximum (minimum) total weight. The weighted matching problem can be solved by a combinatorial algorithm that uses the unweighted Edmonds's algorithm as a subroutine.[2] Kolmogorov provides an efficient C++ implementation of this.[7]
References
- ^ Edmonds, Jack (1965). "Paths, trees, and flowers". Canad. J. Math. 17: 449–467. doi:10.4153/CJM-1965-045-4.
- ^ a b Lovász, László; Plummer, Michael (1986). Matching Theory. Akadémiai Kiadó. ISBN 9630541688.
- ^ a b Karp, Richard, "Edmonds's Non-Bipartite Matching Algorithm", Course Notes. U. C. Berkeley, http://www.cs.berkeley.edu/~karp/greatalgo/lecture05.pdf
- ^ a b Tarjan, Robert, "Sketchy Notes on Edmonds' Incredible Shrinking Blossom Algorithm for General Matching", Course Notes, Department of Computer Science, Princeton University, http://www.cs.dartmouth.edu/~ac/Teach/CS105-Winter05/Handouts/tarjan-blossom.pdf
- ^ Kenyon, Claire; Lovász, László, "Algorithmic Discrete Mathematics", Technical Report CS-TR-251-90, Department of Computer Science, Princeton University
- ^ Micali, Silvio; Vazirani, Vijay (1980), "An O(V1/2E) algorithm for finding maximum matching in general graphs", 21st Annual Symposium on Foundations of Computer Science, (IEEE Computer Society Press, New York): 17–27
- ^ Kolmogorov, Vladimir (2009), "Blossom V: A new implementation of a minimum cost perfect matching algorithm", Mathematical Programming Computation 1 (1): 43–67, http://www.cs.ucl.ac.uk/staff/V.Kolmogorov/papers/BLOSSOM5.html